Martin Laurberg is a Structural Biologist working in the field of Protein Synthesis. He is a research associate in the laboratory of Prof. Harry F. Noller at the University of California at Santa Cruz.
Protein Synthesis
Protein Synthesis is the biological link between genotype and phenotype, and the process is crucial in maintaining life. Without new protein, the cell would die. The ribosome is the biomolecular machine facilitating protein synthesis from transcribed genes. Two thirds of all currently known antibiotics target the ribosome, and ribosomal research therefore plays a central role in modern healthcare.
Structural studies of the ribosome, e.g. by X-ray Crystallography, can decipher in atomic detail, how this ancient molecular machine performs some of the most essential molecular tasks in biology. Crystallographic three-dimensional 'snapshots' of each of the ribosome`s many operational states, reveal its dynamics and enzymatic chemistry, how antibiotics `throw a spanner in the works` and how novel antibiotics may be designed.
The crystal structure above explains how peptide release factor RF1 (yellow) decodes a STOP-codon (UAA) of the mRNA (green), and in turn catalyzes the hydrolytic release of the synthesized protein (not shown) attached at the 3' end of the P-site tRNA (orange) in a concluding step of protein synthesis. The structure delivers the long-sought completing piece of information to the genetic code, explaining the structural basis for which STOP codons are decoded by class I release factors, while SENSE codons are read by basepairing tRNAs.
The Ribosome is an Ancient Biomolecular Machine
Life on Earth is based on six chemical elements, H, C, N, O, S and P (and perhaps As). They form biological compounds, such as the nucleotide building blocks of DNA and RNA and the amino acids of proteins.
Proteins and DNA may have emerged as catalytic and genomic improvements to an ancient, now extinct, 'RNA World'. The ribosome is essentially an RNA-based machine, possibly as a result of its molecular heridity. At the evolutionary onset of the Domains of Life from the Last Universal Ancestor, the ribosome was already well in place and remains similar at its structural core in all organisms to this day, despite (huge) structural variation elsewhere pertaining to divergent evolution. Therefore we share fundamentally similar ribosomes with all other current species and e.g. the self-confident T.rex roaming the land of the Cretaceous Period (~65 mio years ago) and the first cyanobacteria quietly loafing the oceans of the Archean Eon, ~2.5 billion years ago.
Class I ribosomal release factors in Bacteria and Eukarya (and Archaea) are unrelated in sequence and fold (not shown), and may have evolved as molecular mimics of STOP codon specific tRNAs of the last universal ancestor. Indeed, the above crystal structure reveals RF1 as a possible stereochemical tRNA-mimic in the hydrolysis reaction.
Structural Studies of the ribosome bring key insights to evolution, to the origin of life, and perhaps even to our own evolutionary future..
Please visit the Center for Molecular Biology of RNA at the University of California, Santa Cruz
|

Ribosome Crystal

X-ray Diffraction
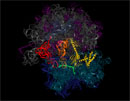
Ribosome Structure
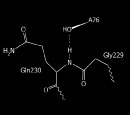
Revealed Chemistry
|







